Combining MRI and digital histopathologic imaging features boosts the accuracy of predictions for overall glioma survival compared to using either of these features alone, according to research published July 23 in Radiology: Imaging Cancer.
The results could help clinicians better care for brain cancer patients, wrote a team led by Saima Rathore, PhD, of the University of Pennsylvania in Philadelphia.
"Although the currently applicable treatment options, which include surgical resection, radiation therapy, chemotherapy, and advanced combination treatment, have expanded during the last couple of decades, the prognosis for patients with high-grade glioma ... remains poor, with disease generally recurring after seemingly successful initial treatment," the team noted. "The survival duration without treatment and the response to treatment vary substantially across individuals, as do the molecular and imaging profiles of the tumors. Hence, there is a need to develop imaging-based markers that are prognostic of patient outcome to potentially assist in building comprehensive and patient-centric treatment plans."
To this end, Rathore's group conducted a study that included multiparametric MRI and histopathologic images from 171 patients with either high-grade or low-grade glioma; data were taken from the Cancer Imaging Archive for the years 1983 to 2008.
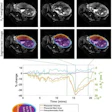
The authors found that patients' overall median survival was 467 days, 350 days for those with high-grade glioma, and 595 days for those with low-grade disease. The team identified 14 MRI and 12 histopathologic imaging features that were predictive for overall survival for patients with both high- and low-grade glioma. It also found that the MRI plus histopathologic image models had a higher overall concordance index compared to either of these imaging datasets alone (0.79 compared with 0.70 for MRI and 0.67 for histopathologic imaging); the combination also performed better for predicting overall survival in both high-grade and low-grade glioma patients, the team found.
| Comparison of predictive models for overall glioma survival | |||
| Concordance index | MRI + histopathologic features | MRI features alone | Histopathologic features alone |
| All patients | 0.79 | 0.70 | 0.67 |
| High-grade glioma patients | 0.78 | 0.68 | 0.64 |
| Low-grade glioma patients | 0.88 | 0.62 | 0.62 |
The study findings confirm other research that has also shown the benefit of combining MRI and histopathologic imaging features to predict outcomes in glioma patients, according to the team.
"[Previous studies have shown] that integrated features improved survival prediction compared with those of clinical and radiographic features alone," the group concluded. "In consensus with these studies, we consistently found that integrating MRI and histopathologic features improved the performance of the prognostic model."




![Overview of the study design. (A) The fully automated deep learning framework was developed to estimate body composition (BC) (defined as subcutaneous adipose tissue [SAT] in liters; visceral adipose tissue [VAT] in liters; skeletal muscle [SM] in liters; SM fat fraction [SMFF] as a percentage; and intramuscular adipose tissue [IMAT] in deciliters) from MRI. The fully automated framework comprised one model (model 1) to quantify different BC measures (SAT, VAT, SM, SMFF, and IMAT) as three-dimensional (3D) measures from whole-body MRI scans. The second model (model 2) was trained to identify standardized anatomic landmarks along the craniocaudal body axis (z coordinate field), which allowed for subdividing the whole-body measures into different subregions typically examined on clinical routine MRI scans (chest, abdomen, and pelvis). (B) BC was quantified from whole-body MRI in over 66,000 individuals from two large population-based cohort studies, the UK Biobank (UKB) (36,317 individuals) and the German National Cohort (NAKO) (30,291 individuals). Bar graphs show age distribution by sex and cohort. BMI = body mass index. (C) After the performance assessment of the fully automated framework, the change in BC measures, distributions, and profiles across age decades were investigated. Age-, sex-, and height-adjusted body composition reference curves were calculated and made publicly available in a web-based z-score calculator (https://circ-ml.github.io).](https://img.auntminnie.com/mindful/smg/workspaces/default/uploads/2026/05/body-comp.XgAjTfPj1W.jpg?auto=format%2Ccompress&fit=crop&h=100&q=70&w=100)




![Overview of the study design. (A) The fully automated deep learning framework was developed to estimate body composition (BC) (defined as subcutaneous adipose tissue [SAT] in liters; visceral adipose tissue [VAT] in liters; skeletal muscle [SM] in liters; SM fat fraction [SMFF] as a percentage; and intramuscular adipose tissue [IMAT] in deciliters) from MRI. The fully automated framework comprised one model (model 1) to quantify different BC measures (SAT, VAT, SM, SMFF, and IMAT) as three-dimensional (3D) measures from whole-body MRI scans. The second model (model 2) was trained to identify standardized anatomic landmarks along the craniocaudal body axis (z coordinate field), which allowed for subdividing the whole-body measures into different subregions typically examined on clinical routine MRI scans (chest, abdomen, and pelvis). (B) BC was quantified from whole-body MRI in over 66,000 individuals from two large population-based cohort studies, the UK Biobank (UKB) (36,317 individuals) and the German National Cohort (NAKO) (30,291 individuals). Bar graphs show age distribution by sex and cohort. BMI = body mass index. (C) After the performance assessment of the fully automated framework, the change in BC measures, distributions, and profiles across age decades were investigated. Age-, sex-, and height-adjusted body composition reference curves were calculated and made publicly available in a web-based z-score calculator (https://circ-ml.github.io).](https://img.auntminnie.com/mindful/smg/workspaces/default/uploads/2026/05/body-comp.XgAjTfPj1W.jpg?auto=format%2Ccompress&fit=crop&h=112&q=70&w=112)