A deep-learning (DL) algorithm that segments 3D pectoralis muscle volume (PMV) shows better reproducibility and stronger associations with chronic obstructive pulmonary disease (COPD) than an analysis that uses 2D pectoralis muscle area (PMA) data, researchers have reported.
A team led by Miranda Kirby, PhD, of Toronto Metropolitan University in Canada, suggested that automated PMV extraction from routine CT scans could help clinicians identify COPD patients with comorbid sarcopenia who may benefit from pulmonary rehabilitation targeting muscle loss. The group's results were published March 5 in Radiology: Cardiothoracic Imaging.
"Compared with the established 2D single-section PMA, the 3D PMV offered greater reproducibility and showed stronger associations with clinical outcomes in individuals with COPD," the investigators noted.
COPD is a multisystem disease that affects organ systems in addition to the lungs, and those with the disease experience chronic low-grade inflammation that can lead to muscle atrophy, or sarcopenia, they explained.
Patients with COPD often undergo CT imaging, and identifying those who also have comorbid sarcopenia could help clinicians set rehabilitation plans to target muscle loss, according to the authors. They investigated the use of a deep-learning algorithm to assess the health of the pectoralis muscle either by volume or by area via a study that included 1,235 participants, of which 634 had COPD and 601 did not.
The team developed a U-Net deep-learning model for PMV segmentation and applied it to information taken from the Canadian Cohort Obstructive Lung Disease (CanCOLD) study, which included data collected from November 2009 to July 2015. It used randomly sampled CT scans from CanCOLD for model training (n = 96), for validation (n = 16), for internal testing (n = 32), and an external dataset for external testing (n = 32). The researchers tracked any differences between patients with or without COPD and associations with forced expiratory volume in one second (FEV1), diffusing capacity of the lungs for carbon monoxide (Dlco), and peak oxygen uptake during exercise (VO2).
The investigators reported the following:
Performance of a deep-learning algorithm that segments pectoralis muscle volume for identifying sarcopenia | |
Type of dataset | Dice similarity coefficient |
| Training and validation | 0.94 |
| Internal testing | 0.93 |
| External testing | 0.92 |
The team also found that both PMA and PMV were reduced in patients with COPD (p < 0.05) and that PMV was more strongly associated with FEV1, Dlco, and VO2 values.
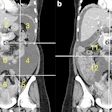
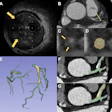
![Images show the pectoralis muscles of a healthy male individual who never smoked (age, 66 years; height, 178 cm; body mass index [BMI, calculated as weight in kilograms divided by height in meters squared], 28.4; number of cigarette pack-years, 0; forced expiratory volume in 1 second [FEV1], 97.6% predicted; FEV1: forced vital capacity [FVC] ratio, 0.71; pectoralis muscle area [PMA], 59.4 cm2; pectoralis muscle volume [PMV], 764 cm3) and a male individual with a smoking history and chronic obstructive pulmonary disorder (COPD) (age, 66 years; height, 178 cm; BMI, 27.5; number of cigarette pack-years, 43.2, FEV1, 48% predicted; FEV1:FVC, 0.56; PMA, 35 cm2; PMV, 480.8 cm3) from the Canadian Cohort Obstructive Lung Disease (i.e., CanCOLD) study. The CT image is shown in the axial plane. The PMV is automatically extracted using the developed deep learning model and overlayed onto the lungs for visual clarity.](https://img.auntminnie.com/mindful/smg/workspaces/default/uploads/2026/03/genkin.25LqljVF0y.jpg?auto=format%2Ccompress&crop=focalpoint&fit=max&q=70&w=400) Images show the pectoralis muscles of a healthy male individual who never smoked (age, 66 years; height, 178 cm; body mass index [BMI, calculated as weight in kilograms divided by height in meters squared], 28.4; number of cigarette pack-years, 0; forced
Images show the pectoralis muscles of a healthy male individual who never smoked (age, 66 years; height, 178 cm; body mass index [BMI, calculated as weight in kilograms divided by height in meters squared], 28.4; number of cigarette pack-years, 0; forced
expiratory volume in 1 second [FEV1], 97.6% predicted; FEV1: forced vital capacity [FVC] ratio, 0.71; pectoralis muscle area [PMA], 59.4
cm2; pectoralis muscle volume [PMV], 764 cm3) and a male individual with a smoking history and chronic obstructive pulmonary disorder (COPD) (age, 66 years; height, 178 cm; BMI, 27.5; number of cigarette pack-years, 43.2, FEV1, 48% predicted; FEV1:FVC, 0.56; PMA, 35 cm2; PMV, 480.8 cm3) from the Canadian Cohort Obstructive Lung Disease (i.e., CanCOLD) study. The CT image is shown in the axial plane. The PMV is automatically extracted using the developed deep learning model and overlayed onto the lungs for visual clarity.RSNA
The research results "encourage the use of the PMV as a surrogate measure of muscle mass in chest CT scans, serving as a potential marker of comorbid sarcopenia in those with COPD," according to the authors.
"Future studies should investigate normative population reference values to determine abnormally low CT-derived PMV sarcopenic thresholds," they concluded.
Access the full study here.